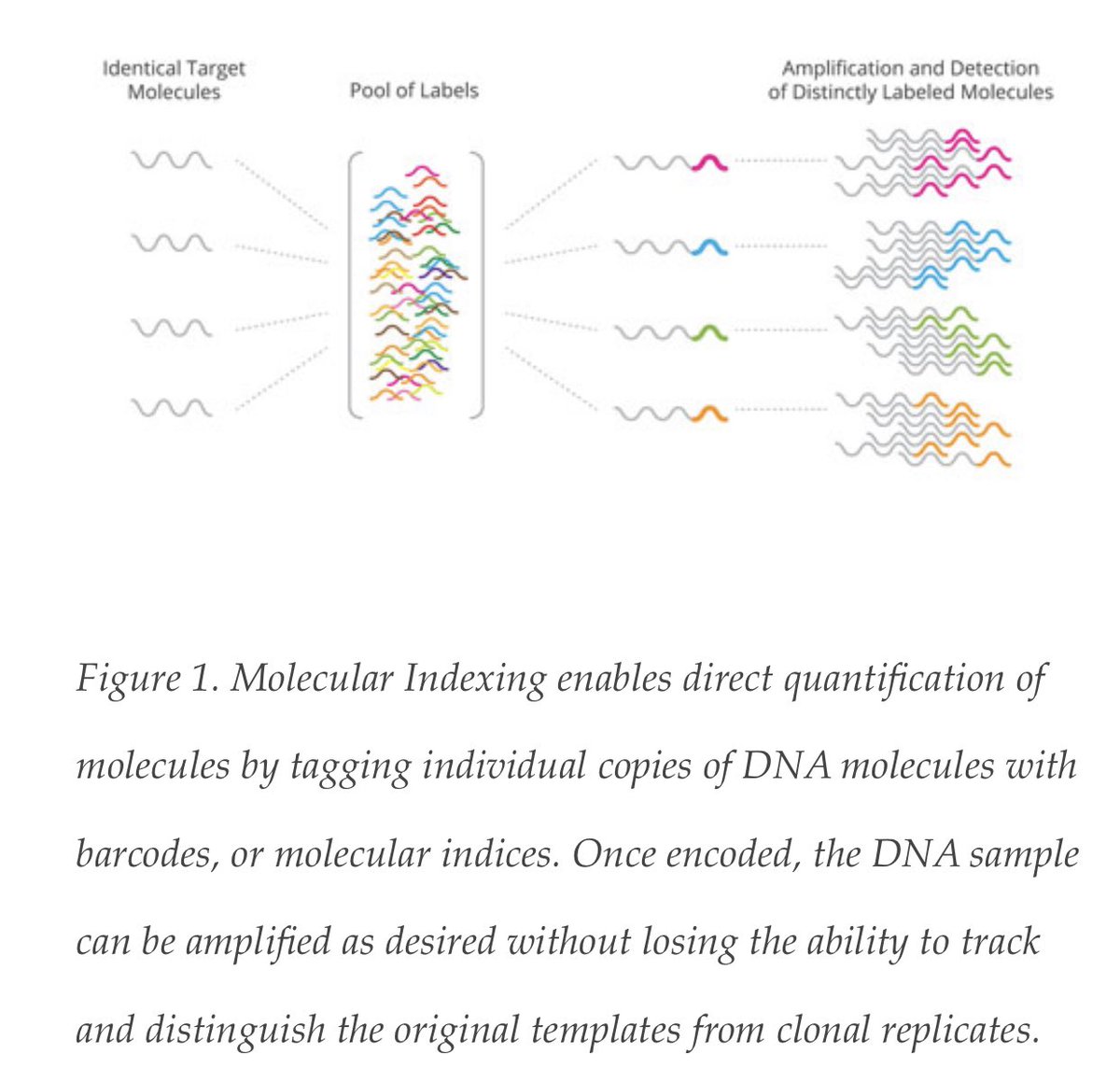
230 days RNA positive post infection.
Turn off the asymptomatic PCR pipeline or we’ll never get out of this.
Turn off the asymptomatic PCR pipeline or we’ll never get out of this.
https://twitter.com/microbeminded2/status/1473721448342687754
Very thorough.
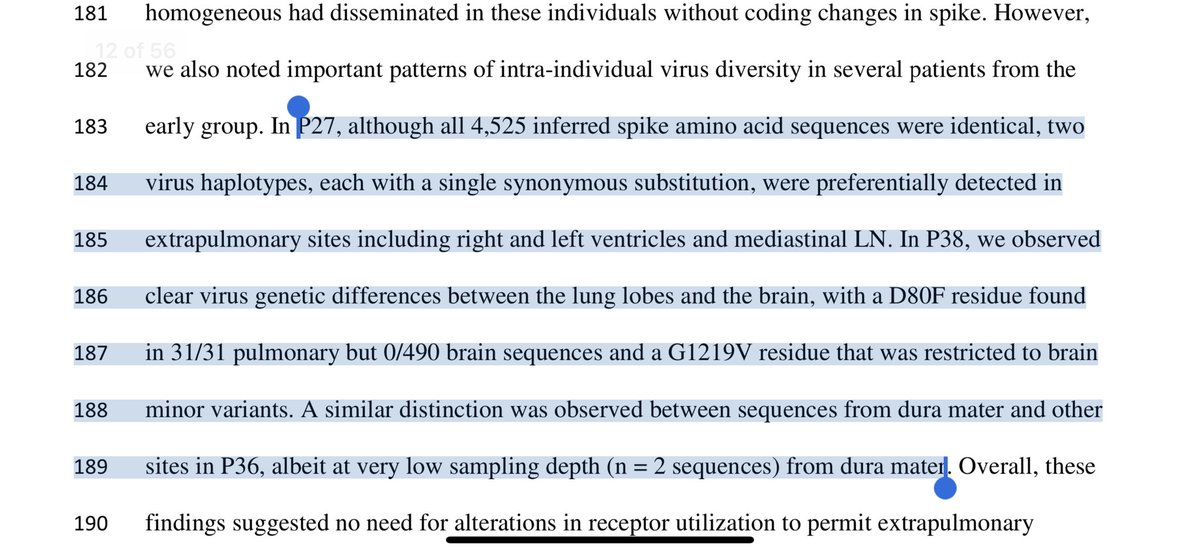
Not only applied a live-dead like PCR looking at sgRNA, they also cultured the virus and looked for immune histochemistry confirmation of proteins.
Not only applied a live-dead like PCR looking at sgRNA, they also cultured the virus and looked for immune histochemistry confirmation of proteins.
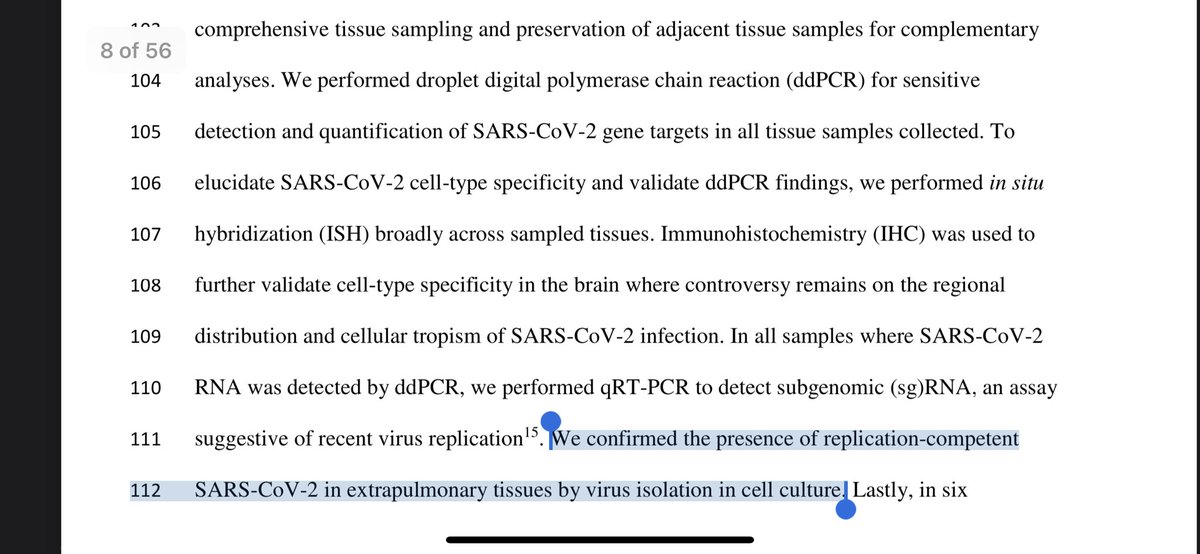

And yes.
One patients duration of illness was 230 days.
Just block the clown on this thread trying to claim this is an RNA measurement from a tissue that’s been dead for 230 days.
Very low signal/noise from that account.
One patients duration of illness was 230 days.
Just block the clown on this thread trying to claim this is an RNA measurement from a tissue that’s been dead for 230 days.
Very low signal/noise from that account.

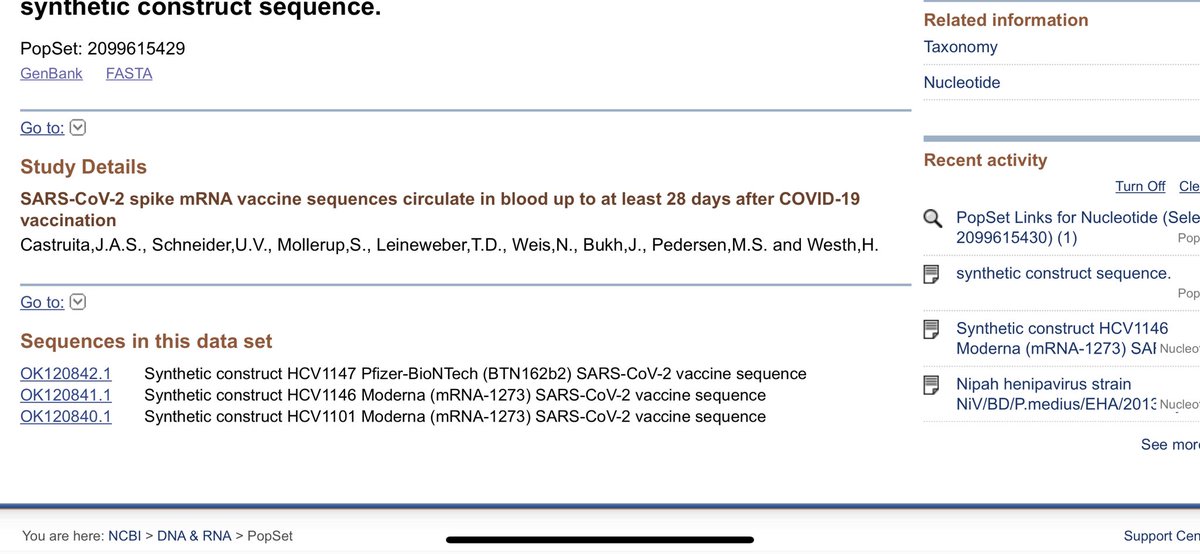
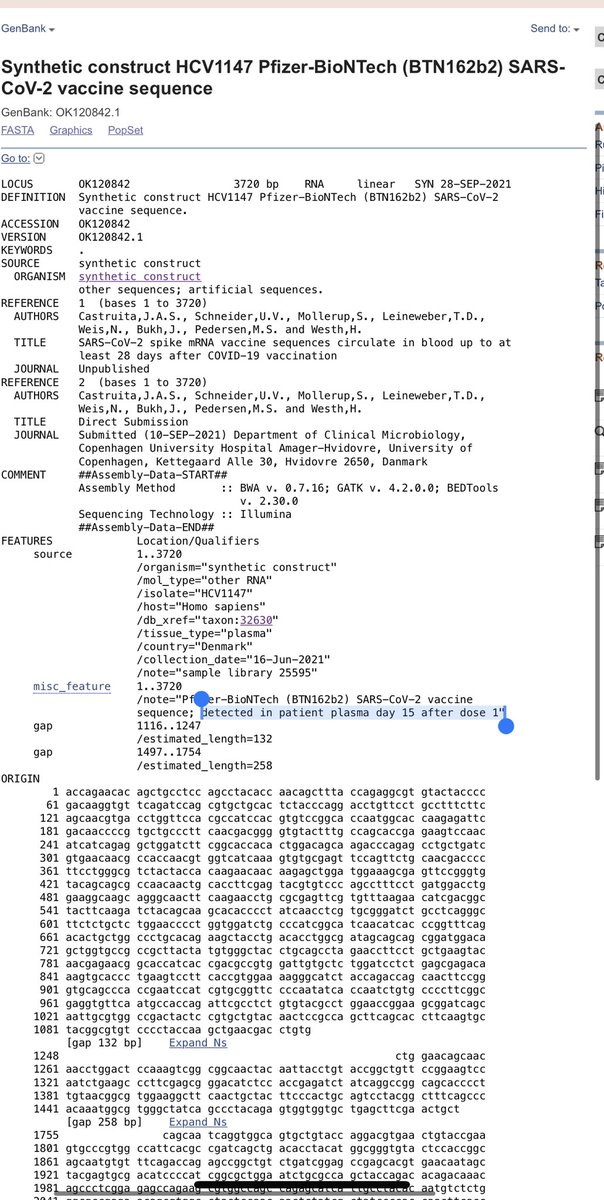
A few folks have asked about the vax status.
They would need a Vax specific PCR process to pick that up due to the codon optimizations that were done on the Vax.
These folks found it 5/15/28 days in the plasma.

They would need a Vax specific PCR process to pick that up due to the codon optimizations that were done on the Vax.
These folks found it 5/15/28 days in the plasma.

• • •
Missing some Tweet in this thread? You can try to
force a refresh