Can we ‘see’ #autism in the brain? And is there a specific brain signature for the condition?
This is what we found by comparing brain connectivity maps across 16 mouse autism models
➡️tinyurl.com/yxfu2gzn
➡️Majestically led 👏👏👏by @Valerio_Zerbi @NCMLabETH
Thread 1/n
This is what we found by comparing brain connectivity maps across 16 mouse autism models
➡️tinyurl.com/yxfu2gzn
➡️Majestically led 👏👏👏by @Valerio_Zerbi @NCMLabETH
Thread 1/n

Autism is diagnosed based on behavior - but is this condition associated with a specific brain signature of circuit dysfunction?
Despite decades of work, brain imaging has failed to identify such signature as we have evidence of under-connectivity... tinyurl.com/y5uxoe3s
2/n
Despite decades of work, brain imaging has failed to identify such signature as we have evidence of under-connectivity... tinyurl.com/y5uxoe3s
2/n

or both (e.g. a mosaic pattern)
tinyurl.com/y5euyn3p
nature.com/articles/mp201…
So the question now is: what is the origin and significance of these heterogeneous findings?
4/n
tinyurl.com/y5euyn3p
nature.com/articles/mp201…
So the question now is: what is the origin and significance of these heterogeneous findings?
4/n

A common (if not prevailing) interpretation of these findings is ➡️there MUST be an autism-specific brain imprint so the fact we have NOT found it YET implicates that brain imaging is a noisy and an unreliable method to map circuit dysfunction in complex brain disorders
5/n
5/n

However, an equally plausible alternative explanation could be that no autism-specific signature of brain dysfunction has been found.....because there is no such thing!
This would actually be consistent with the large etiological heterogeneity of the spectrum...
6/n
This would actually be consistent with the large etiological heterogeneity of the spectrum...
6/n

...and so heterogeneous imaging findings in autism would be a reflection of underlying etiological heterogeneity.
But how can we rigorously test this hypothesis? Clinical studies are complicated by genetic heterogeneity
and confounding environmental contributions
7/n
But how can we rigorously test this hypothesis? Clinical studies are complicated by genetic heterogeneity
and confounding environmental contributions
7/n

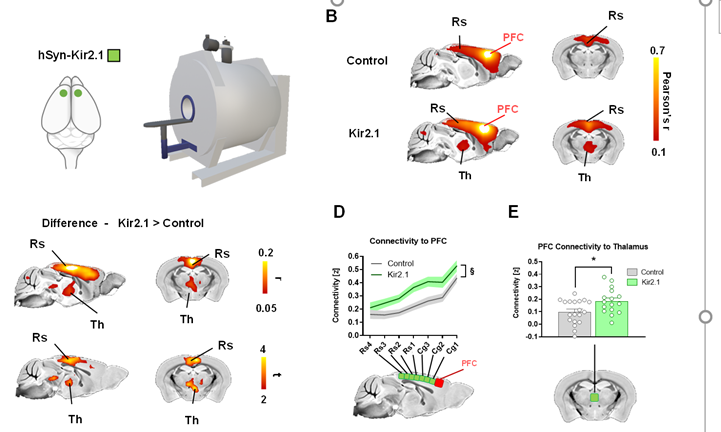
So we turned to mouse🐁 imaging
➡️we established a bi-center study to model the etiological "spectrum" in the 🐁collecting #16 mutations/etiologies
➡️we next mapped and compared the corresponding functional connectivity signature of each mutations using rsfMRI
8/n
➡️we established a bi-center study to model the etiological "spectrum" in the 🐁collecting #16 mutations/etiologies
➡️we next mapped and compared the corresponding functional connectivity signature of each mutations using rsfMRI
8/n

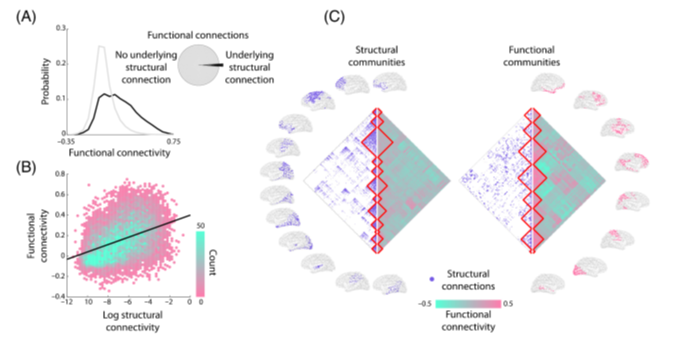
We found that
➡️Most individual scans can be clustered together etiology-wise: so rsfMRI is not so noisy after all..
➡️Autism-related etiologies define a pseudo-continuous connectivity landscape - this argues AGAINST a *unique* signature of circuit dysfunction for autism
9/n
➡️Most individual scans can be clustered together etiology-wise: so rsfMRI is not so noisy after all..
➡️Autism-related etiologies define a pseudo-continuous connectivity landscape - this argues AGAINST a *unique* signature of circuit dysfunction for autism
9/n

What differentiates these connectivity signatures? And can we leverage this heterogeneity to identify cross-etiological connectivity subtypes?
To addressed these question we used unsupervised clustering of connectivity across etiologies
10/n
To addressed these question we used unsupervised clustering of connectivity across etiologies
10/n

and found enlightening results:
➡️No single dysconnectivity signature but n=4 cross-etiological connectivity subtypes with anatomically-specific patterns of dysconnectivity!
➡️some of these subtypes (i.e. #1 & #4) show *opposing* patterns of under- vs- over-connectivity!
11/n
➡️No single dysconnectivity signature but n=4 cross-etiological connectivity subtypes with anatomically-specific patterns of dysconnectivity!
➡️some of these subtypes (i.e. #1 & #4) show *opposing* patterns of under- vs- over-connectivity!
11/n

Taken together our results suggest that
➡️Etiological variability is a *key* determinant of connectivity heterogeneity in autism
➡️Diverging connectivity patterns may explain & reconcile conflicting findings in clinical studies
12/n
➡️Etiological variability is a *key* determinant of connectivity heterogeneity in autism
➡️Diverging connectivity patterns may explain & reconcile conflicting findings in clinical studies
12/n
➡️Last but not least: etiologically-relevant connectivity subtypes could help parse autism heterogeneity
Check-out recent example here⬇️
⚠️Note we did not emphasize role of grouped etiologies here as N=16 is too low to speculate about pathways⚠️
13/n
Check-out recent example here⬇️
https://twitter.com/Gozzi_Ale/status/1314465108136005633
⚠️Note we did not emphasize role of grouped etiologies here as N=16 is too low to speculate about pathways⚠️
13/n
So that's all. Many thanks & kudos to @Valerio_Zerbi for promoting and flawlessly executing the project @MarcoPagani1985 for crucial input + all the stellar collaborators for sharing mouse lines and helping with interpretation @SFARIorg & all funding agencies involved
• • •
Missing some Tweet in this thread? You can try to
force a refresh