We have completely mapped #SARS_CoV_2 mutations that escape binding by LY-CoV555 (antibody that forms the basis for Eli Lilly's bamlanivimab) both alone & in cocktail with LY-CoV016 in a new study led by @tylernstarr w/ @AllieGreaney & @AdamDingens: biorxiv.org/content/10.110… (1/n)
We corroborate recent work showing LY-CoV555 and its cocktail with LY-CoV016 is escaped by mutations in B.1.351 and P.1 viral lineages (E484K and K417N/T, respectively), and also show that LY-CoV555 is affected by the L452R mutation in B.1.429. (2/n)
Specifically, we used complete mapping approach we had previously applied to antibodies in REGN-COV2 (science.sciencemag.org/content/371/65…) to also determine how all RBD mutations affect LY-CoV555 binding. Below are maps of how mutations affect binding (big letter = escape from binding) (3/n) 

Consistent w recent work by Wang et al (David Ho's group, biorxiv.org/content/10.110…), E484K escapes LY-CoV555 and K417N escapes LY-CoV016, which means cocktail won't work against B.1.351. Also, we show L452R affects LY-CoV555 binding. (4/n) 

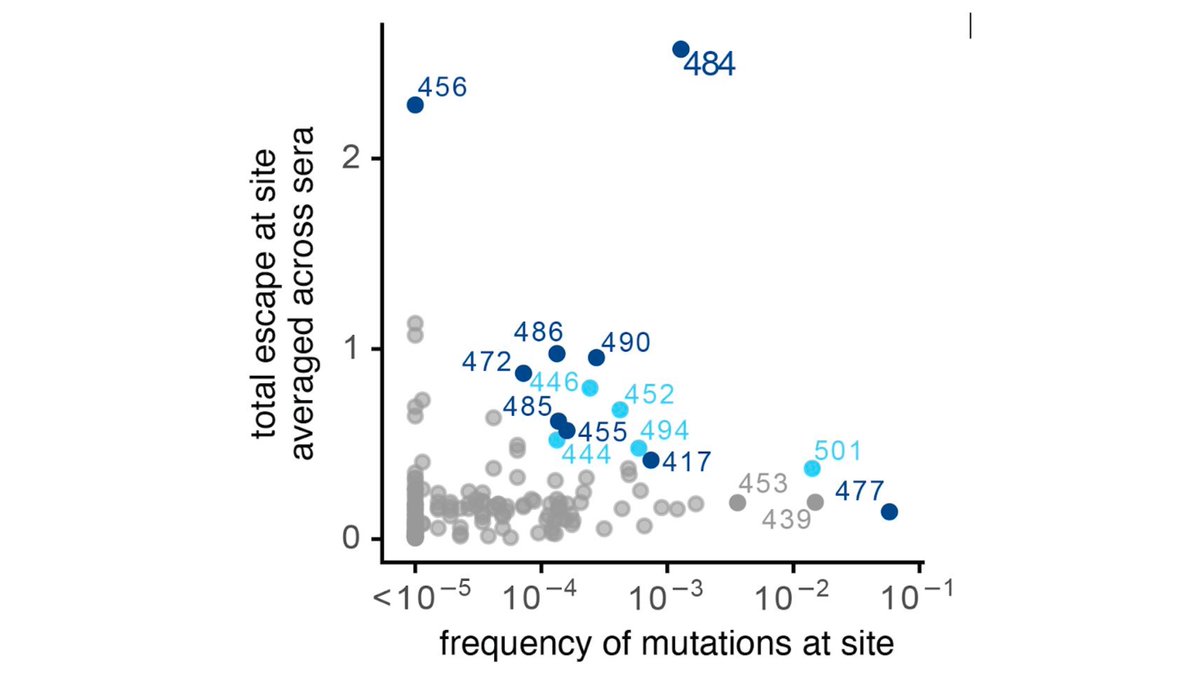
More generally, our maps enable prospective surveillance of other mutations that affect binding by these antibodies. Here are maps of mutations that affect binding by each (y-axis) versus current frequency in sequenced isolates (x-axis). (5/n) 

You can interactively explore our escape maps and download raw data here: jbloomlab.github.io/SARS-CoV-2-RBD… (6/n)
Finally, our results and other recent work in this area strongly suggest a priority should be to diversify epitopes targeted by antibodies in clinical development, such that the same mutations selected under immune pressure don't also escape so many clinical antibodies. (7/n)
• • •
Missing some Tweet in this thread? You can try to
force a refresh