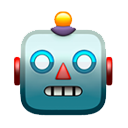
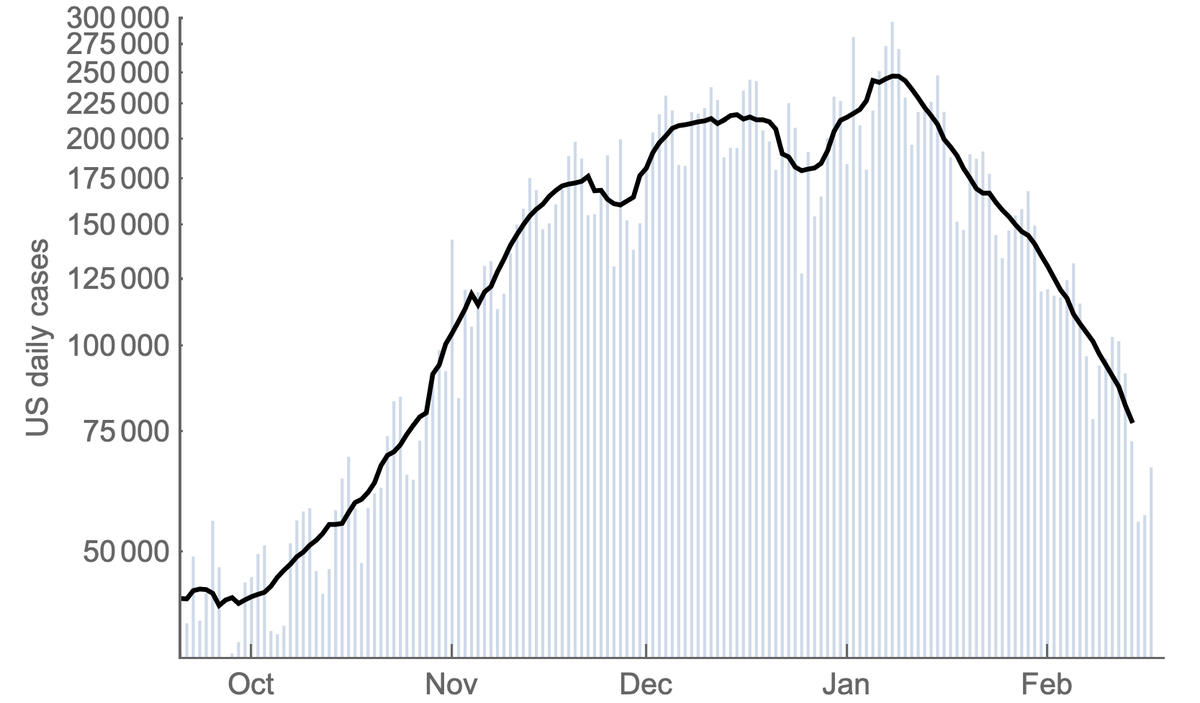
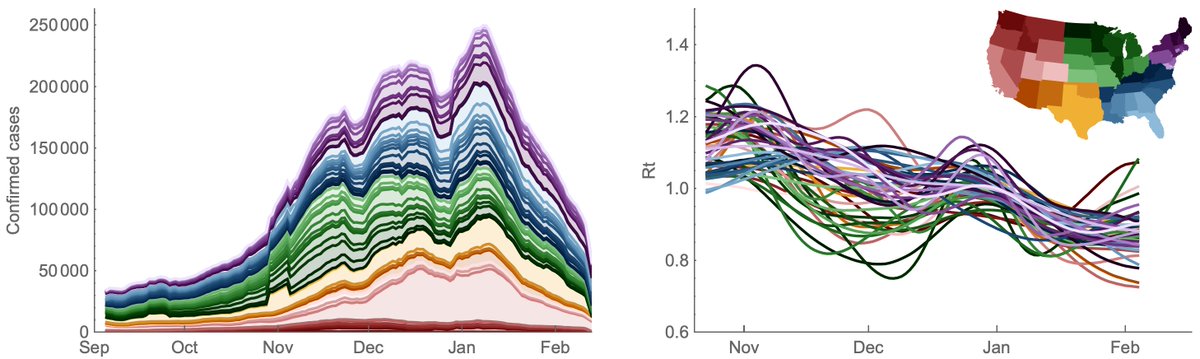
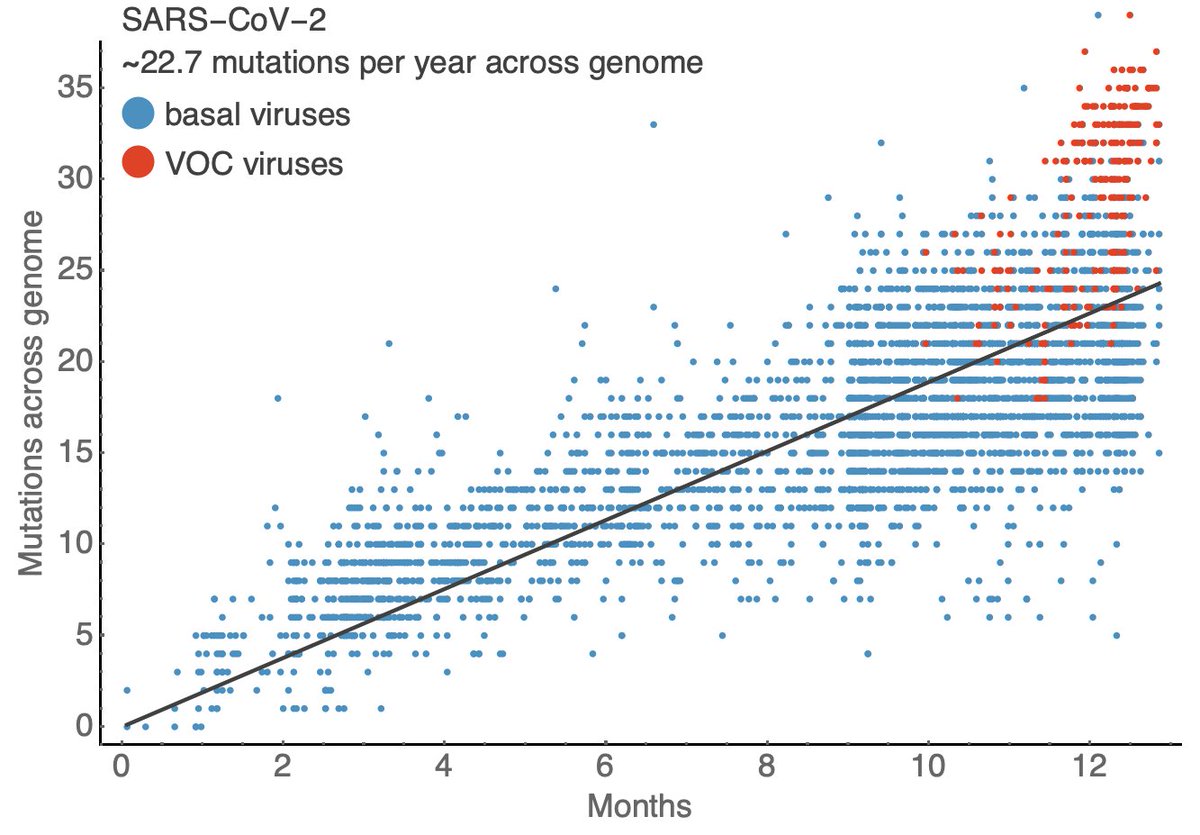
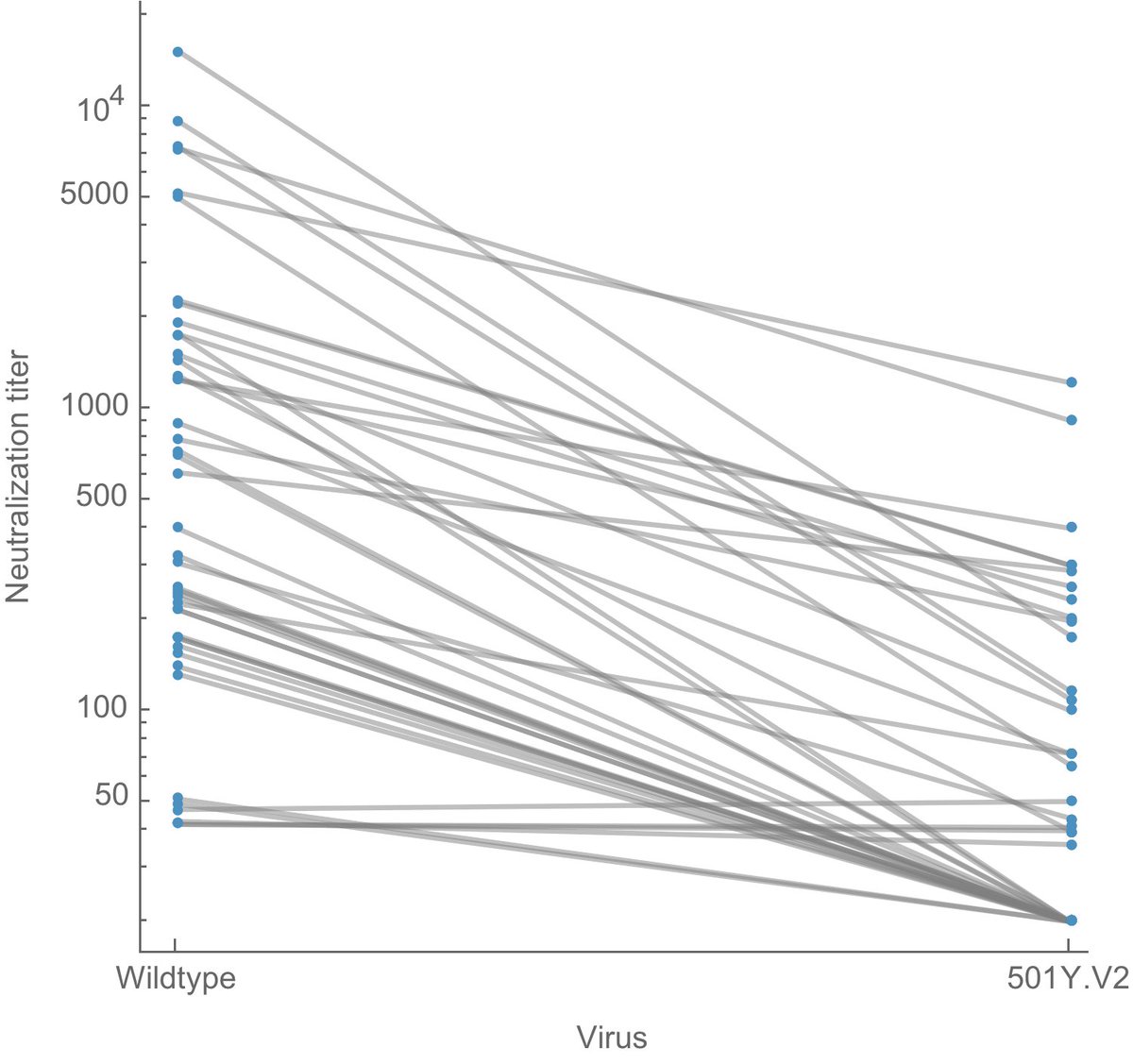
It's hard for me to infer the degree to which new variants are driving the surge in cases in India, but we are seeing rapid growth in frequency of multiple viral variants. 1/5
Here is a @nextstrain view of @GISAID data that focuses on viruses from India and highlights emerging lineages B.1.1.7 (in blue), B.1.351 (in green) and B.1.617 (in orange). Interactive version at nextstrain.org/ncov/asia?c=em…. 2/5 

We can fit a logistic growth model to the full genomic dataset from India for these three lineages, where we see logistic growth as "linear on a logit scale". Each of these lineages is estimated to have similar logistic growth rates of ~0.3 per week. 3/5 
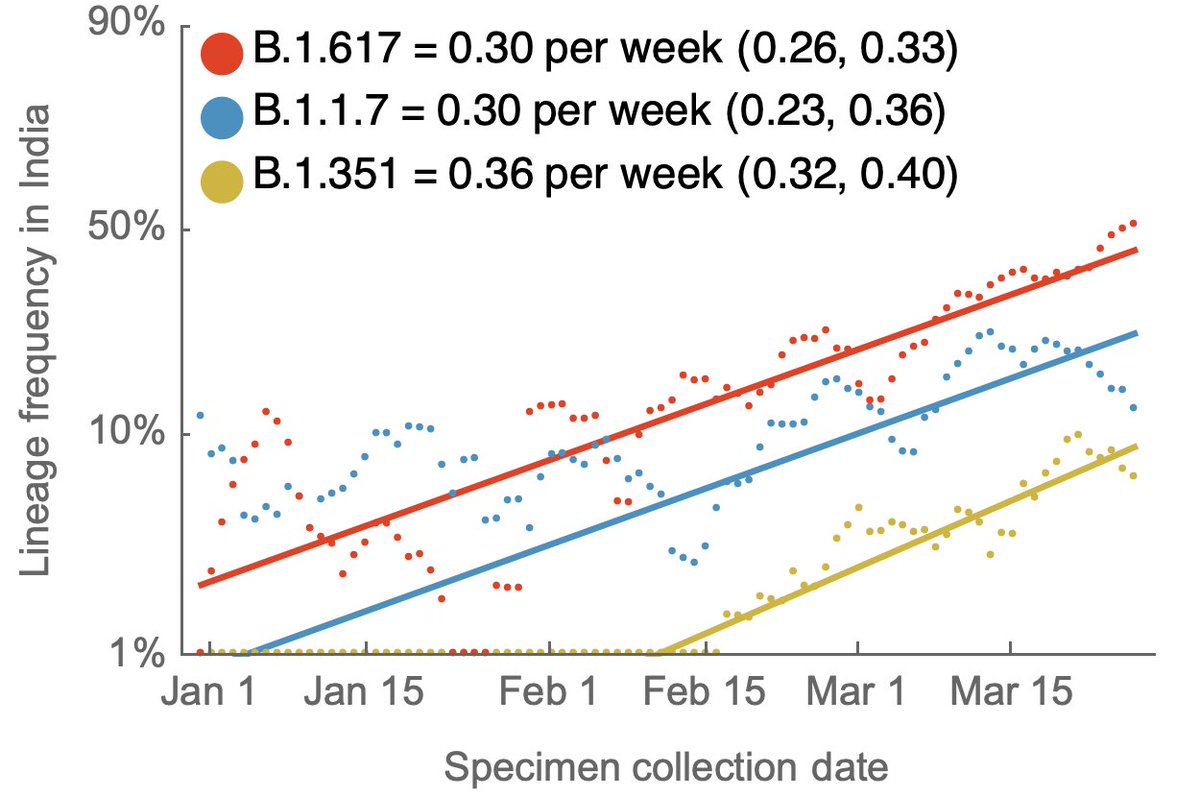
It appears however that B.1.617 was at a higher starting point in January and so frequency at the end of March (we lack data for April) is highest for B.1.617 at around ~45% frequency. 4/5 
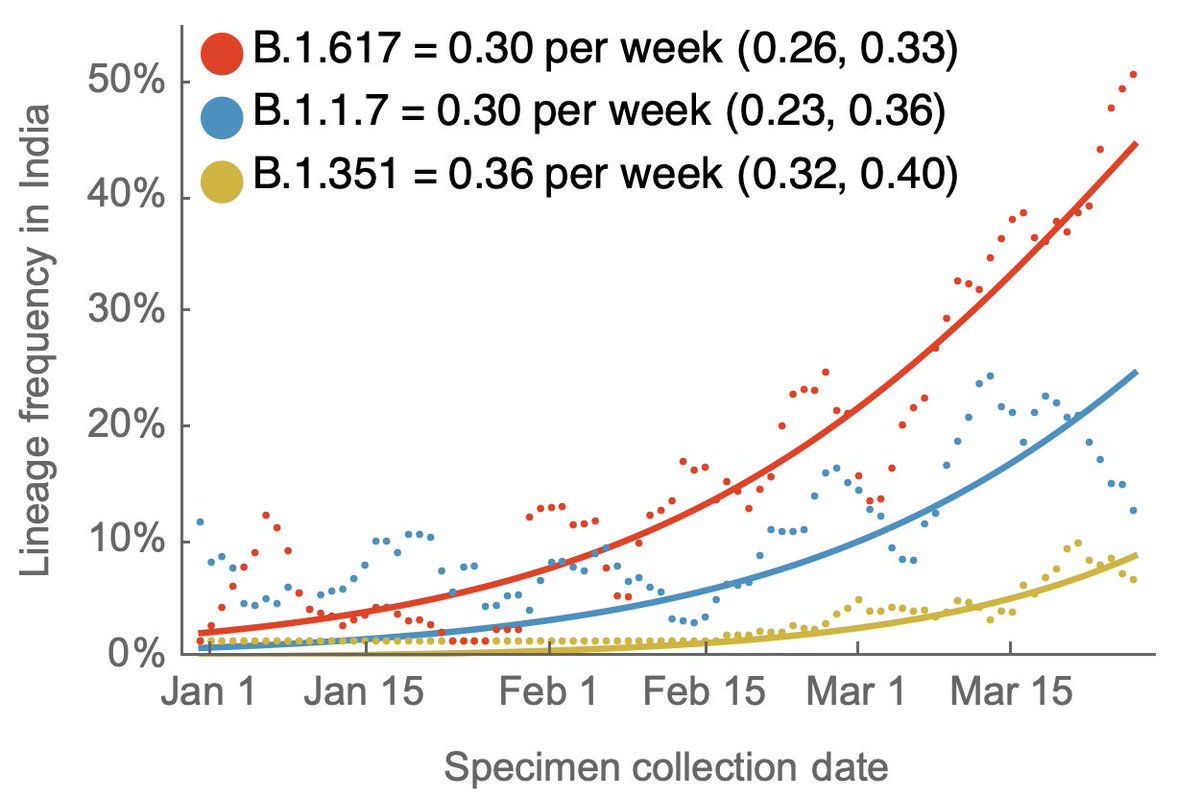
As both B.1.617 and B.1.1.7 further increase in frequency they will be more directly in competition and it should become clear which lineage has the evolutionary advantage. 5/5
• • •
Missing some Tweet in this thread? You can try to
force a refresh