The drivers of the #COVID19 epidemic in India are certainly multifactorial, but we have now seen the viral lineage B.1.617 linked to this epidemic continue to increase in frequency in India and spread rapidly outside of the country. 1/10
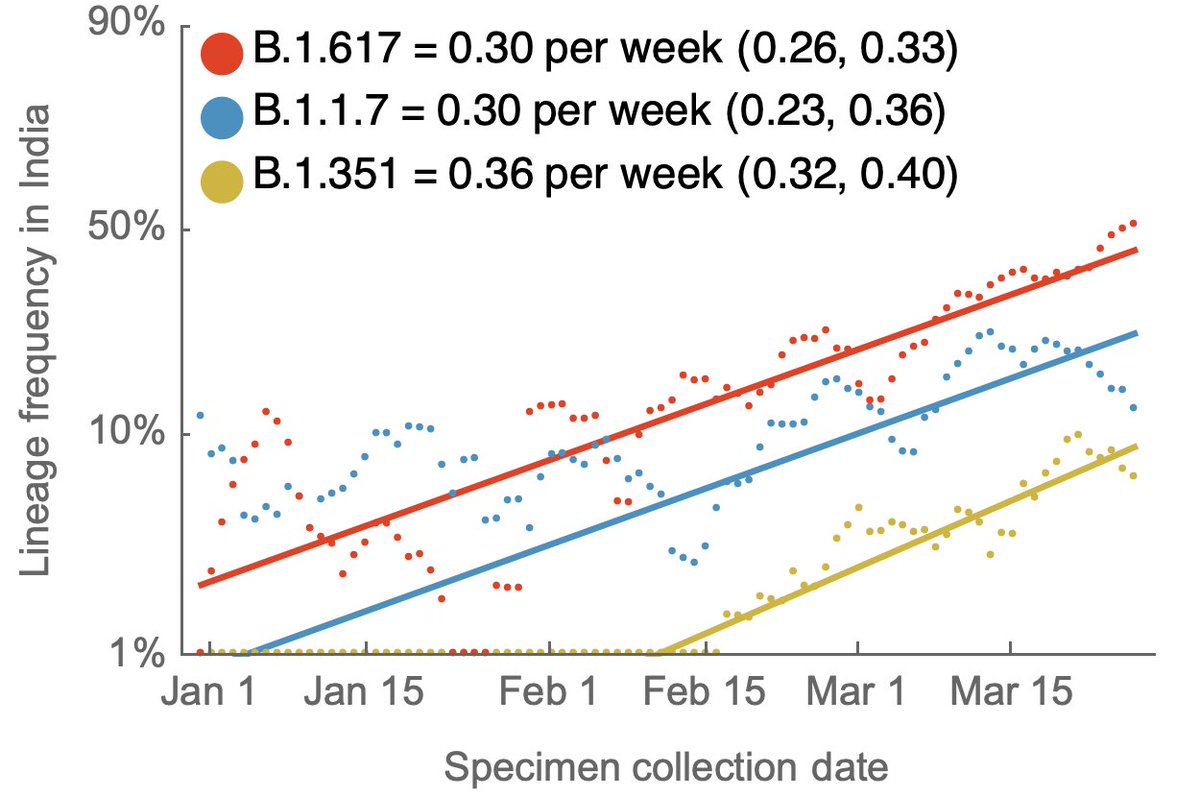
Looking within India there are three primary viral lineages of consequence: B.1.1.7 (in blue) and B.1.351 (in green) introduced into India repeatedly from outside the country and B.1.617 (in yellow) emerging endogenously from within India (nextstrain.org/ncov/asia?c=em…). 2/10 
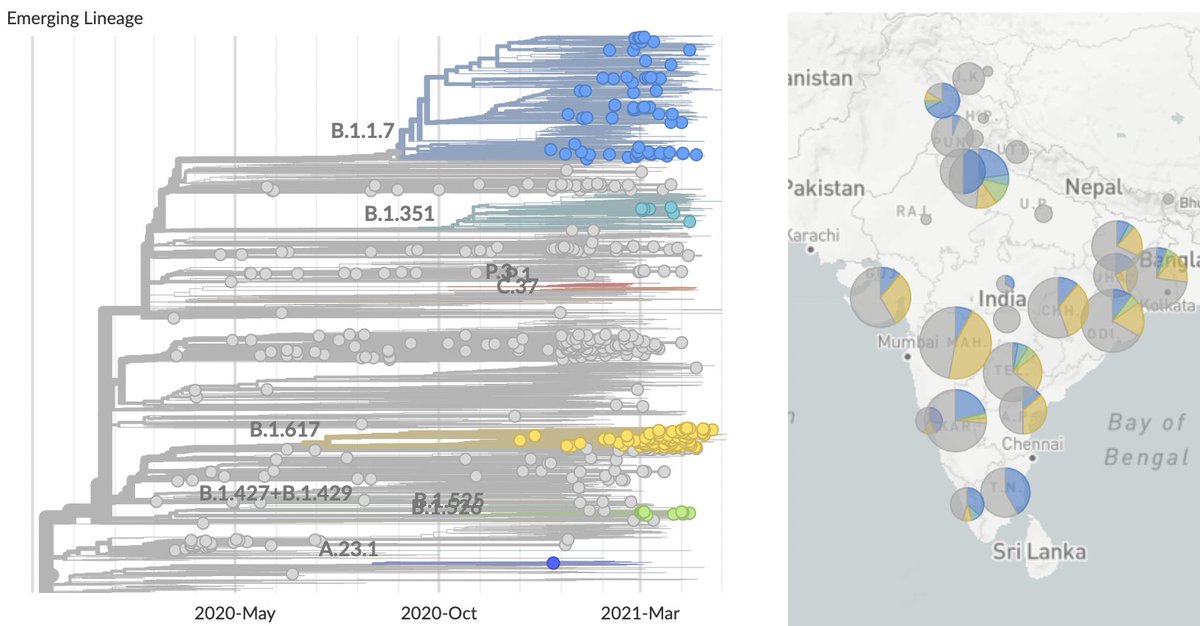
Tracking frequencies over time in sequence data shared to @gisaid shows a continued increase in B.1.617, while recent weeks have shown a decline in B.1.1.7. 3/10 
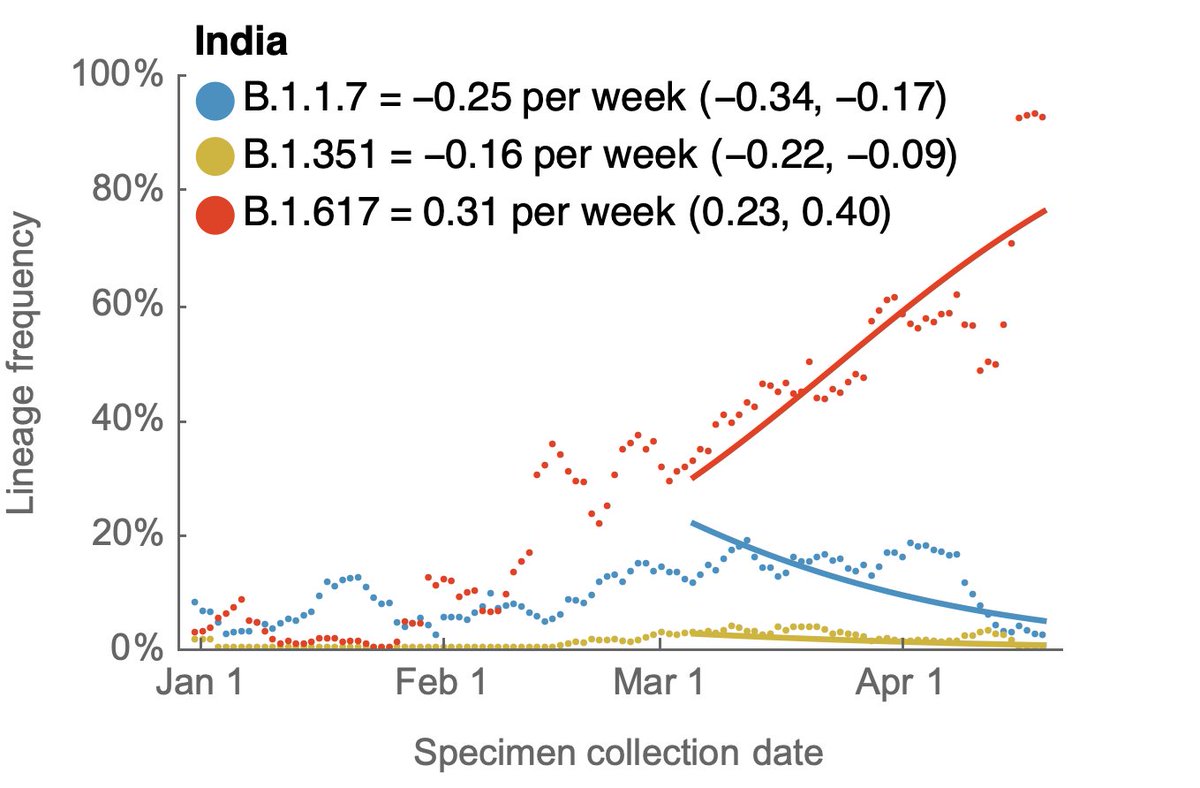
Frequency estimates for April are noisy and end on April 22 due to data availability. There are 883 sequenced genomes for April, but are not strictly in proportion to cases and have a particular geographic focus of Gujarat, Karnataka, Maharashtra and West Bengal. 4/10
Unlike B.1.1.7 and P.1 where mutations comprising the new variant all appeared more-or-less in tandem, the B.1.617 lineage has shown continued evolution since its emergence in mid-2020. 5/10
Basal mutations L452R and P681R exist across the lineage, while sub-lineage B.1.617.1 possesses additional mutations E154K and E484Q and sub-lineage B.1.617.2 possesses additional mutation T478K and commonly 157-158del. 6/10 
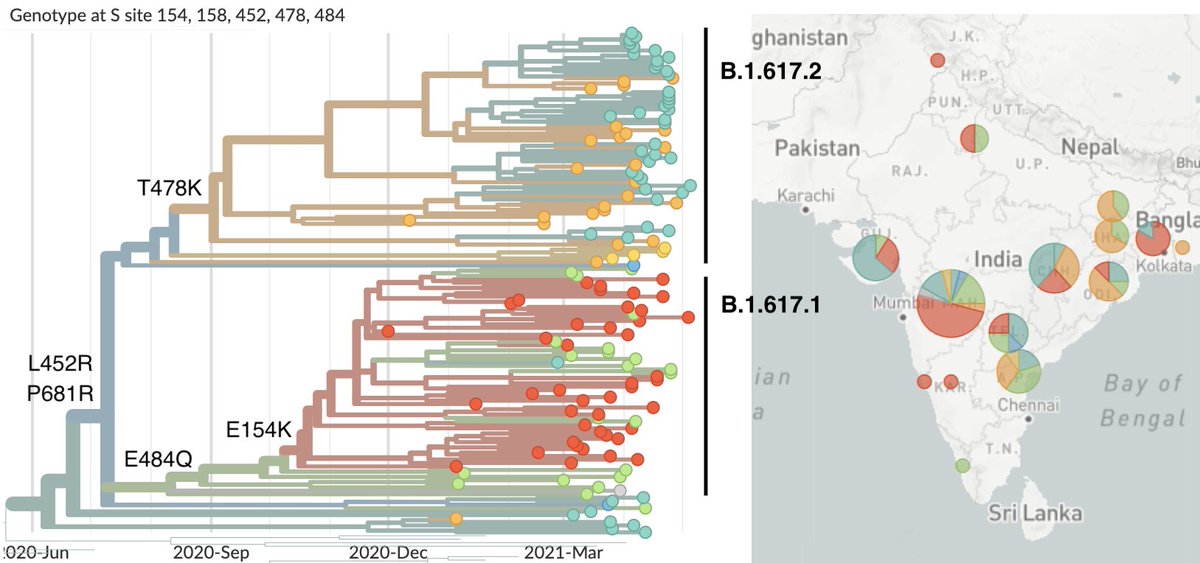
Looking at frequencies of sub-lineages of B.1.617 shows that the overall increase of B.1.617 is driven by sub-lineage B.1.617.2. Here, we see B.1.617.1 declining at a similar pace as B.1.1.7. 7/10 
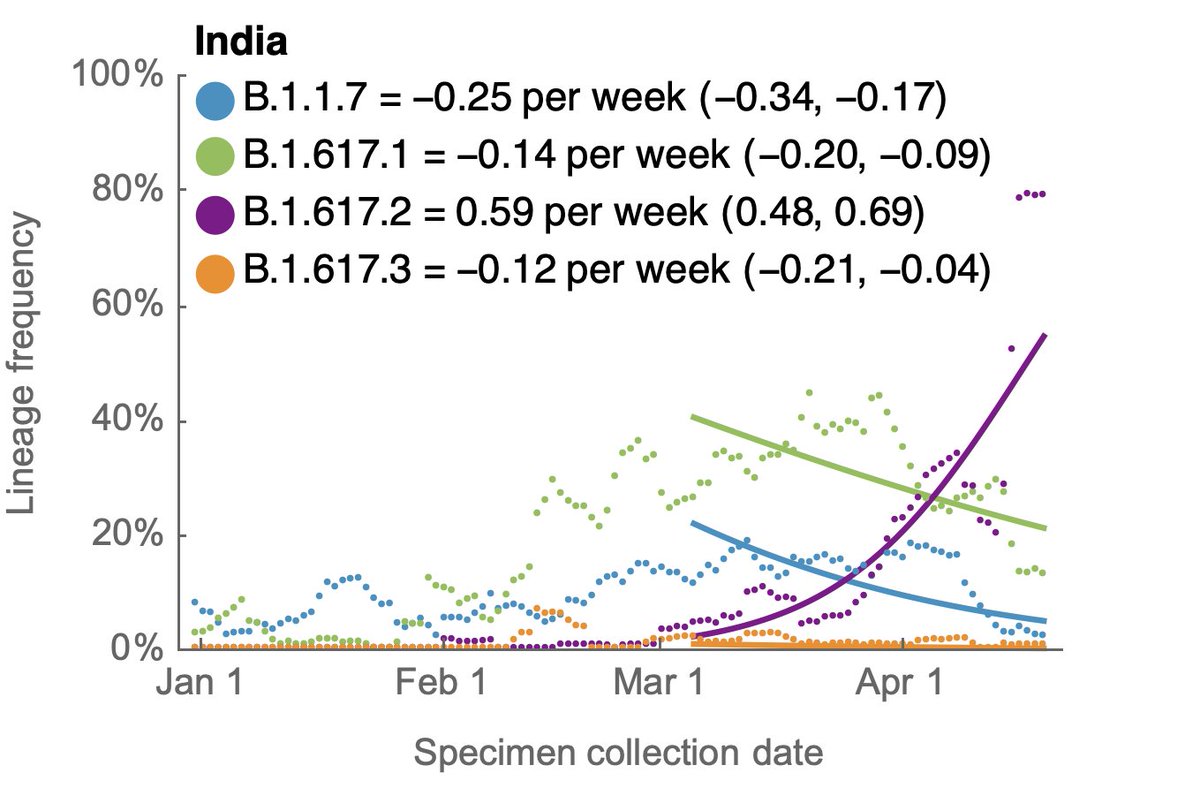
If we look more broadly, B.1.617 has started to spread beyond India and we observe that among clades in the @nextstrain globally subsampled view, B.1.617 is now the most rapidly expanding clade as measured by logistic growth rate (nextstrain.org/ncov/global?br…). 8/10 

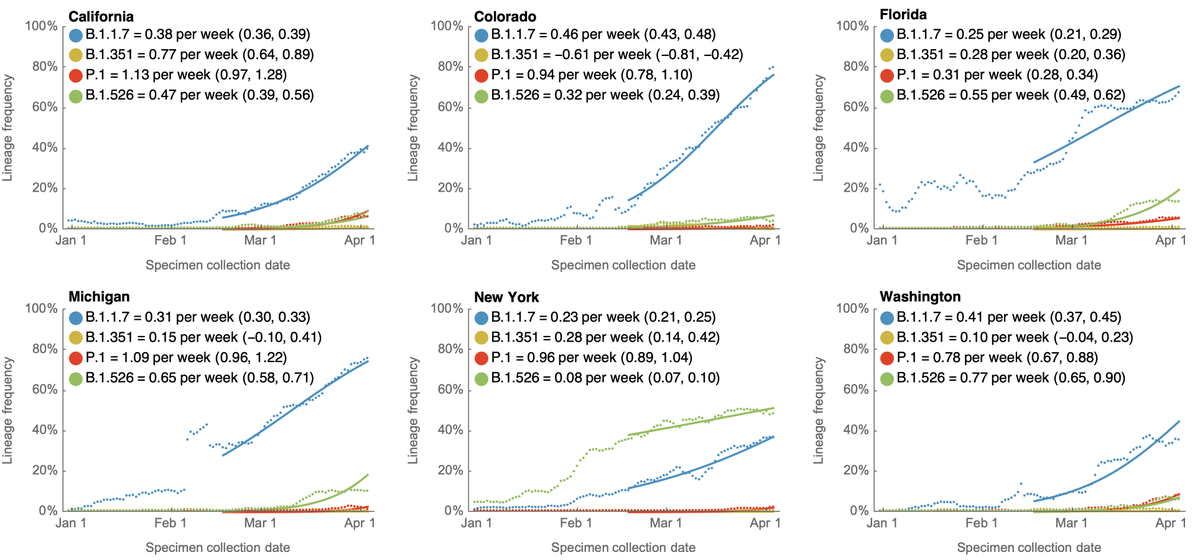
This spread can also be seen in frequency trajectories in individual countries. This is particularly striking in the UK where B.1.617.2 was just announced as a variant of concern by @PHE_uk. 9/10 
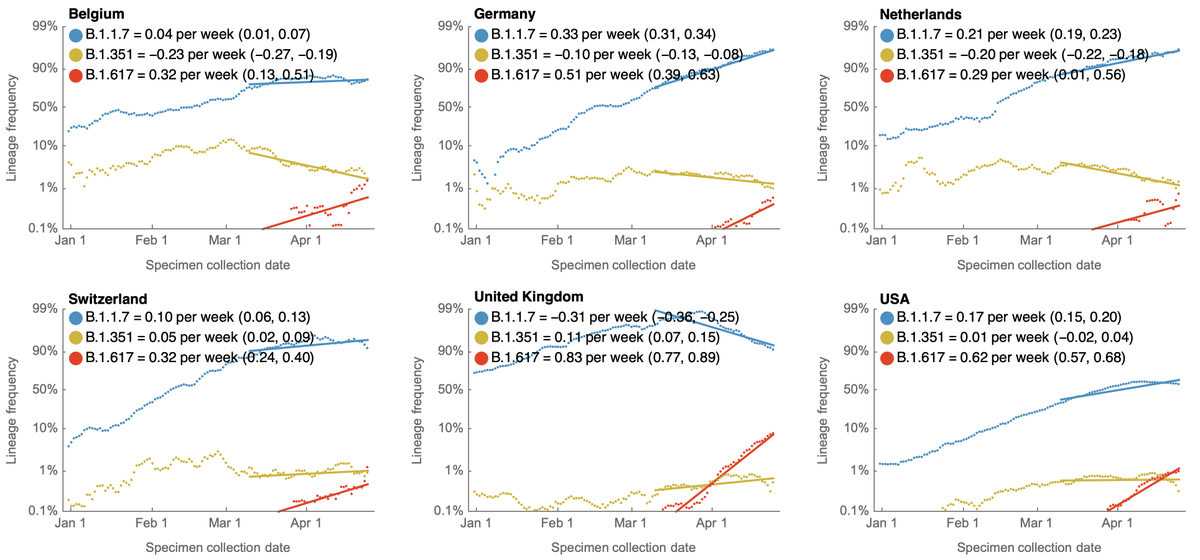
The observed rapid growth of this (sub)-lineage in India and elsewhere would suggest that this virus is potentially highly transmissible. If faster growth than B.1.1.7 in India and in the UK is conclusive, it would suggest that this lineage will spread widely. 10/10
• • •
Missing some Tweet in this thread? You can try to
force a refresh