🔍spatial.libd.org/spatialLIBD/
👩🏾💻github.com/LieberInstitut…
📚research.libd.org/spatialLIBD/
📦github.com/LieberInstitut…
biorxiv.org/content/10.110…
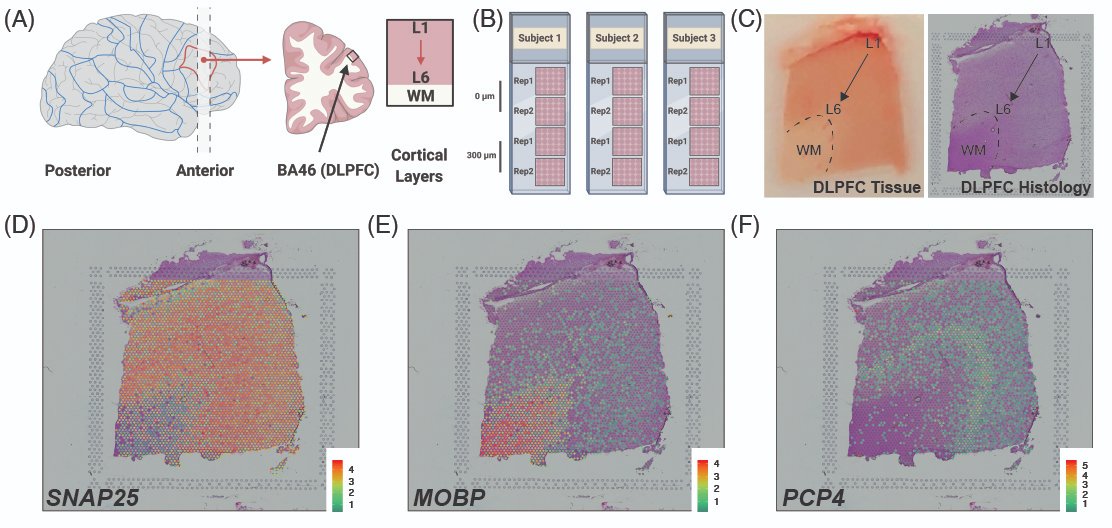
* Largest so far
* Harder to cluster than the example mouse data
* Data is all public
* @Bioconductor #rstats📦
* #shiny web app
2/7 #spatialLIBD
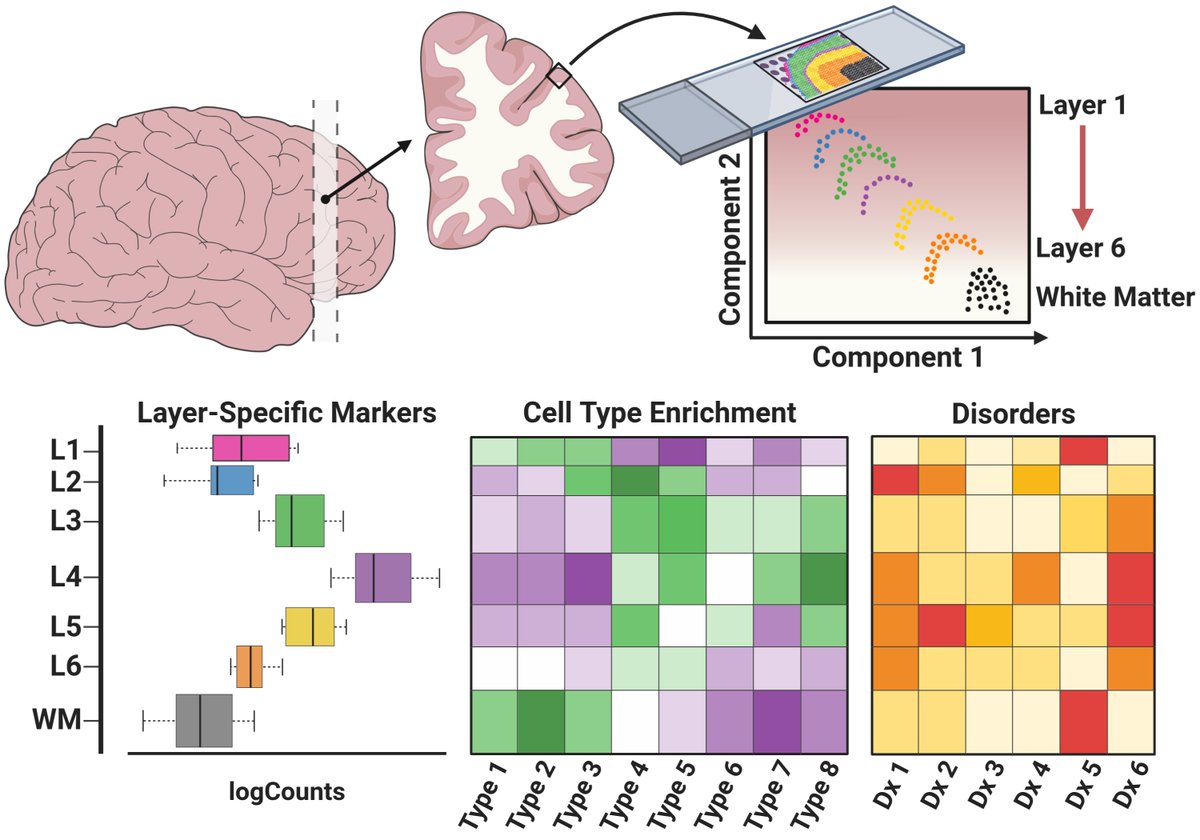
* Identification of layer-specific markers
* smFISH validation✅
* spatial registration of snRNA-seq data
* Layer-enrichment of neurodevelopmental and neuropsychiatric gene sets
* Data-driven layer-enriched clustering in the DLPFC
3/7 #spatialLIBD
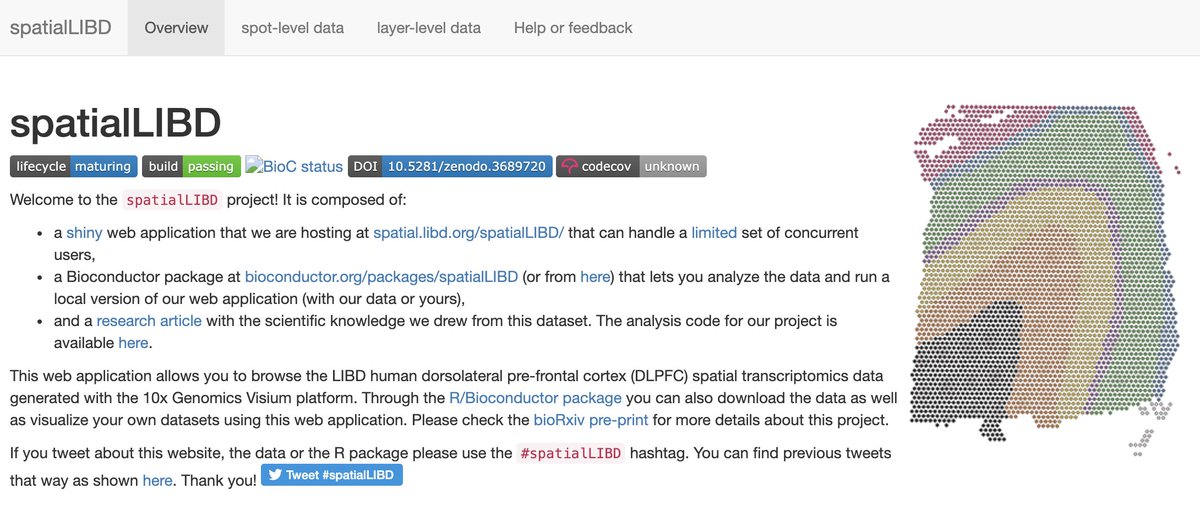
It was a pleasure to be involved in this project!
4/7 #spatialLIBD
research.libd.org/spatialLIBD/
5/7 #spatialLIBD #rstats #Bioconductor #OpenScience
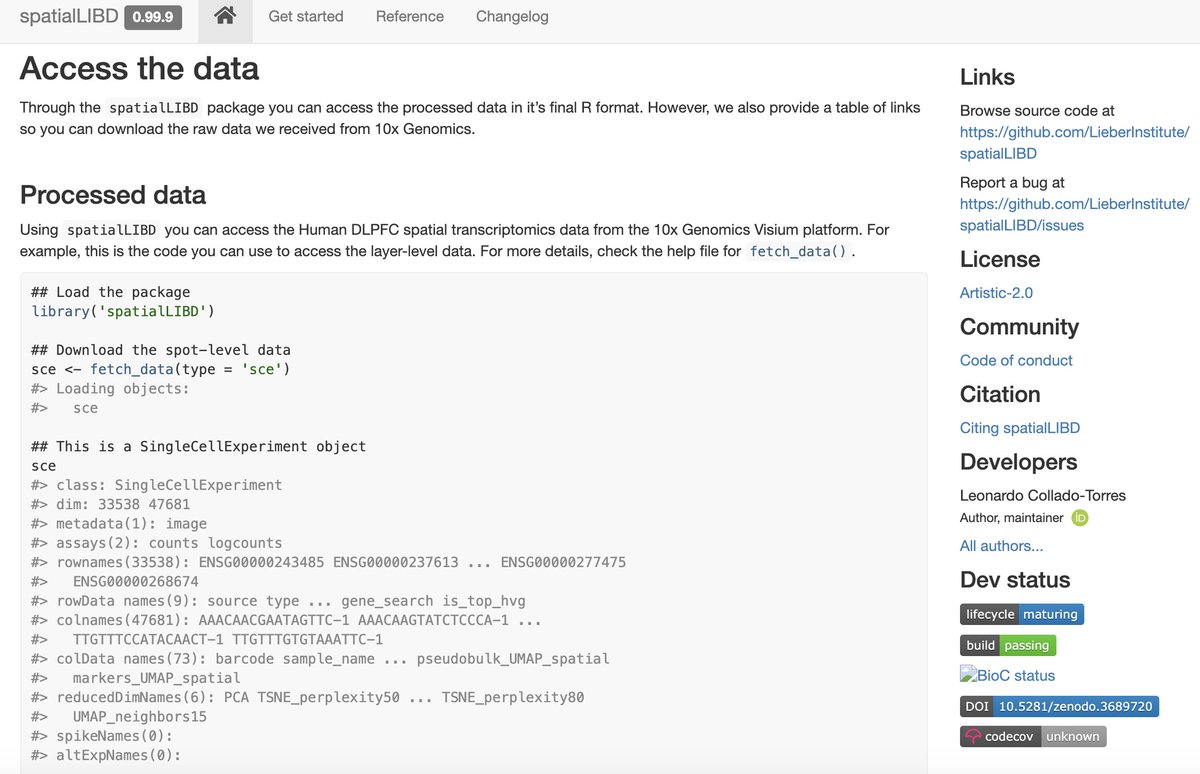
March
19 the-scientist.com
26-27
30 @LIIGH_UNAM
2020 Single Cell Symp.
#BoG2020 @cshlmeetings
@erum2020_conf
#BioC2020 @Bioconductor
6/7 #spatialLIBD

"Diving together into the unknown world of spatial transcriptomics"
lcolladotor.github.io/2020/02/29/div…
7/7 #spatialLIBD #stats #scicomm #IntoTheUnknown