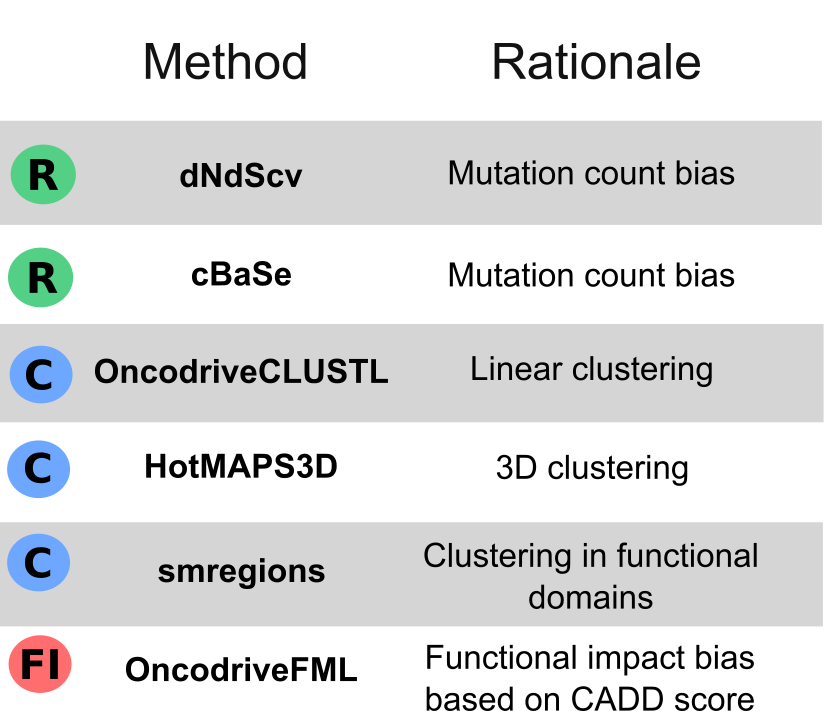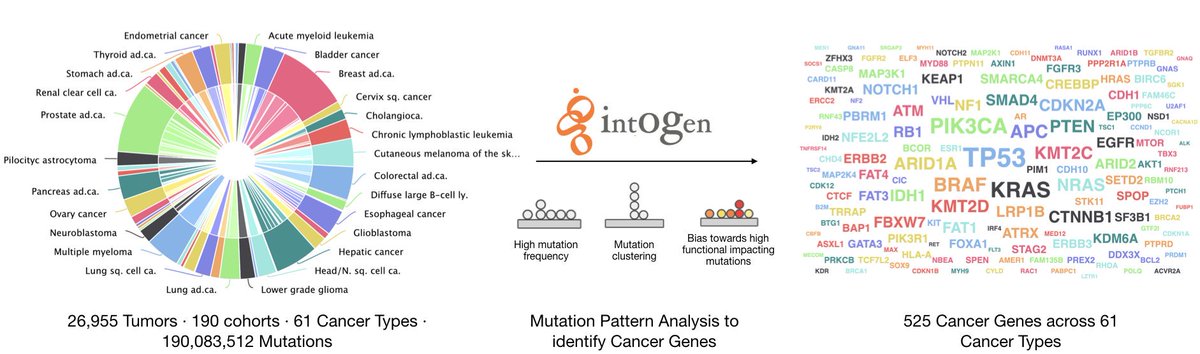
We manually downloaded, processed and annotated tumor genomic data from different sources, such as @cbioportal #PedcBioPortal @icgc_dcc #TCGA #PCAWG @HartwigMedical #TARGET @StJude and others. 190 cohorts in total👇
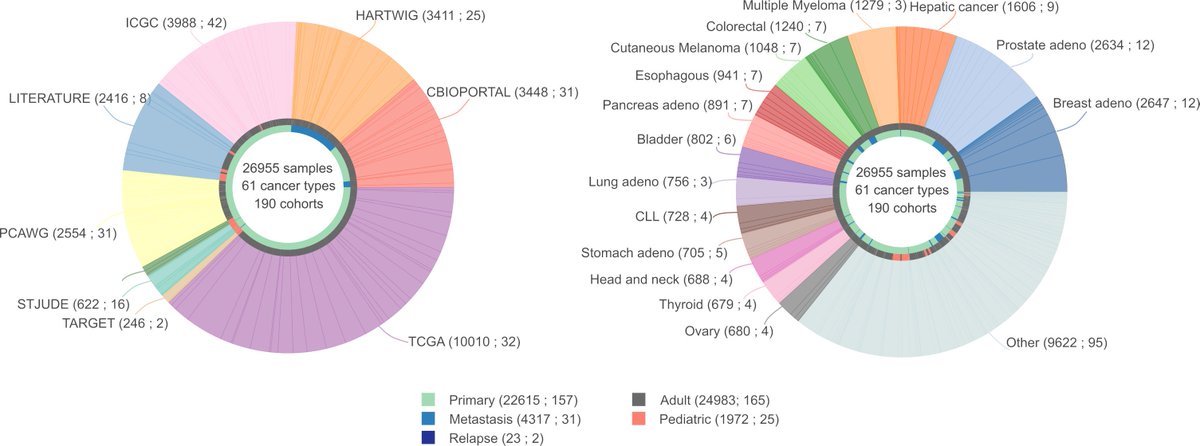
You can search for a gene and see if it has driver mutations in any of the 61 cancer types analyzed. Or you can search for a cancer type and see all genes with driver mutations.
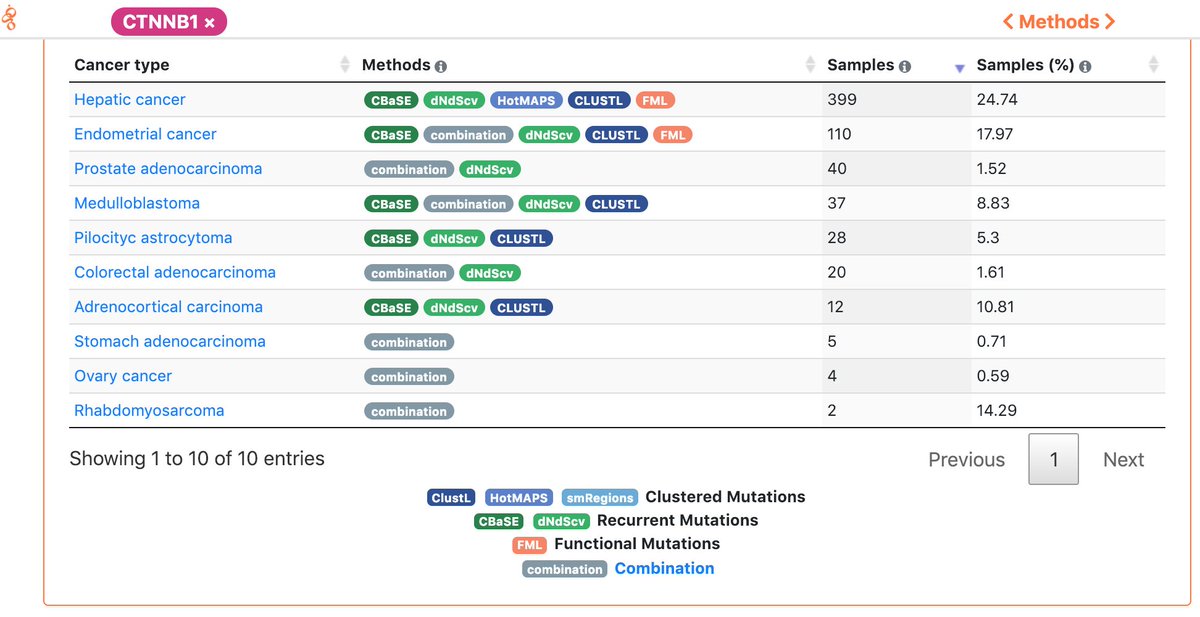

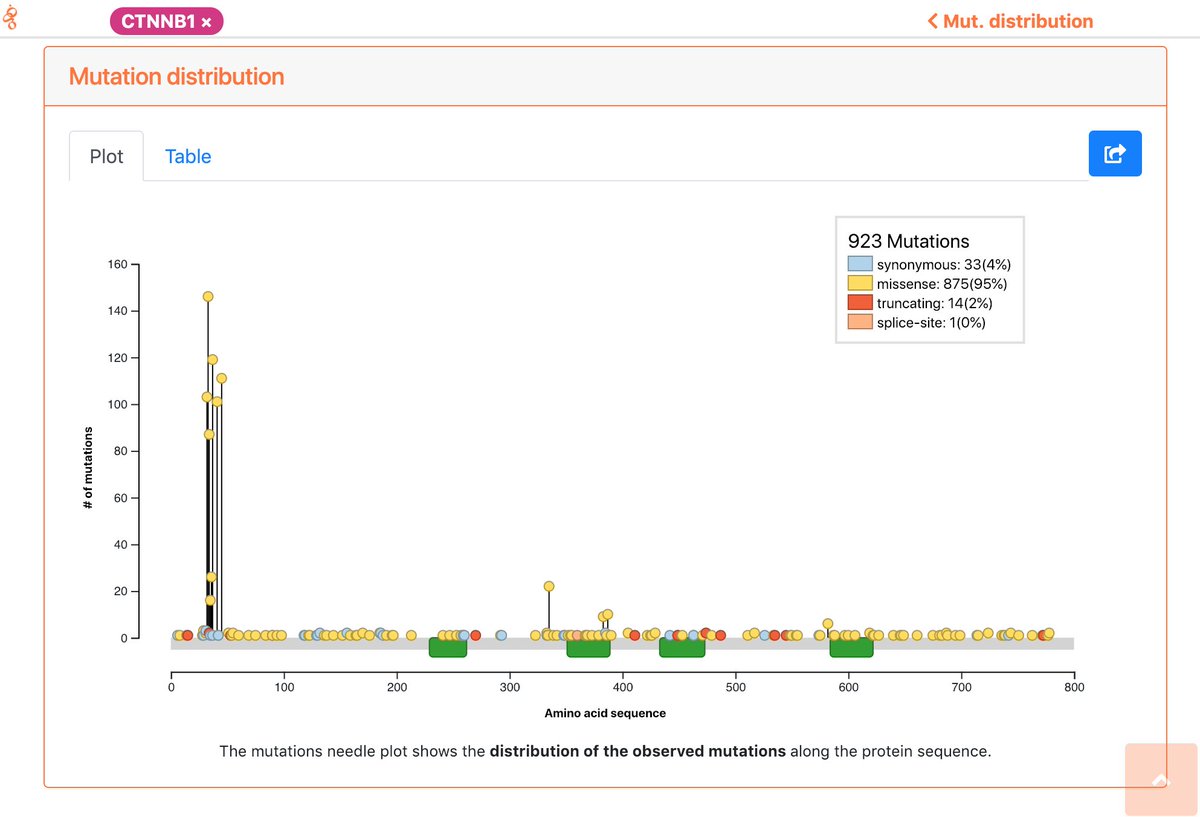
Yes, all the data generated by intOGen can be downloaded and is available for use free of restrictions under the @creativecommons Zero Public Domain Dedication (creativecommons.org/publicdomain/z…).
Yes, we have implemented a @nextflowio pipeline that can be run locally. The source code of the intOGen pipeline is provided under The GNU General Public Licen v3.0
👉Pipeline code: bitbucket.org/intogen/intoge…
👉Documentation: intogen.readthedocs.io/en/latest/data…
IntOGen is a team effort from @bbglab.
See intogen.org/beta/about


We hope to make a convincing case of the benefit of accessible data to progress in science