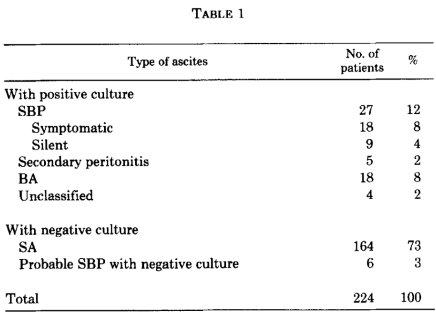
Tentatively calling this #tweetorial "Antibiotic testing: why and how and huh?"
Important caveat: this is a HUGE topic. I’m mostly going to focus on 1) basic rationale and 2) laboratory methods. And 3) try not to say anything wrong/misleading!
All are legit answers! But it's very situation dependent and you'd have different reasons for each. Let’s go through them.
#tweetorial #pathtweet #asmclinmicro

Patient has Enterococcus (of some species) in their blood: I want to treat with vancomycin. Ergo, I should try to grow the bacteria in the presence of vancomycin.
#asmclinmicro
Vancomycin resistant Enterococcus (VRE) is a serious hospital acquired infection (HAI). Which is a pretty big deal (PBD).
#tweetorial #pathtweet #asmclinmicro
Enterococcus species against vanc can trigger an investigation into a possible VRE outbreak in a hospital setting. cdc.gov/hai/organisms/…
#tweetorial #pathtweet #asmclinmicro
There are several different species of rapidly growing mycobacterium.
Sometimes conventional identification methods (e.g. MALDI) can have some trouble calling them to the species level. #tweetorial
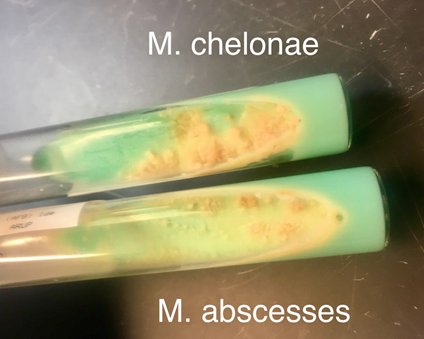
For instance, an isolate gets called as M. fortuitum.
But upon testing it looks resistant to cefoxitin and trimethoprim-sulfa and susceptible to clarithromycin. ...
This looks like M. immunogenum’s resistance pattern. They should re-test the identification.
#tweetorial #asmclinmicro
Characterizing the antibiotic susceptibility profiles of unusual bugs can be very helpful! ... IF you get the information out there.
You may turn to PubMed and see if something’s published.
If someone’s tested it before against a battery of antibiotics, that can help you know what to start with.
But for the second part of this #tweetorial: how do you actually DO this?
#AntibioticResistance
Small paper disk w/ antibiotics in it. Streak agar plate with a known concentration of bacteria. Put disks down. The closer to the disks the bacteria grows = the more resistant to the antibiotic effect it is.
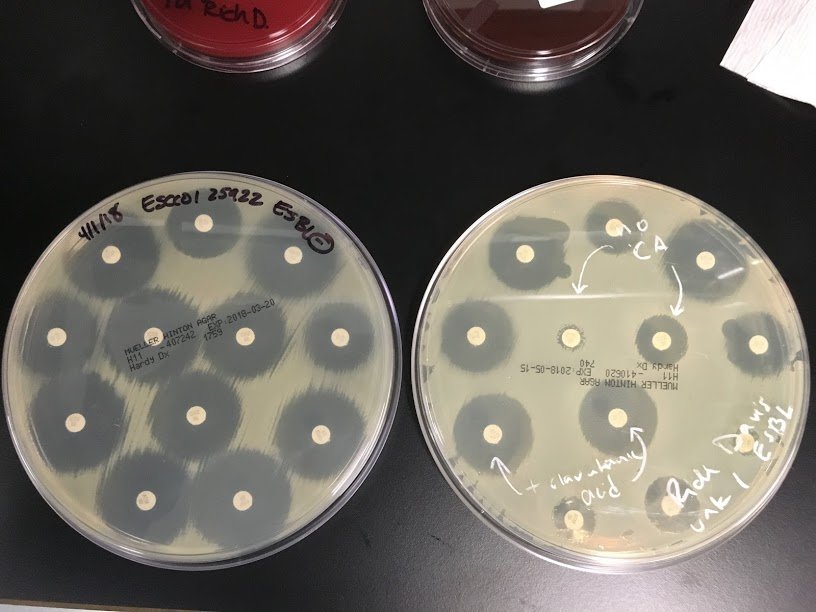
Antibotics are put into a well in smaller dilutions. Put known concentration of bacteria into the wells. Look to see the LOWEST concentration where no bacteria grows. More resistant = growth at higher concentrations.

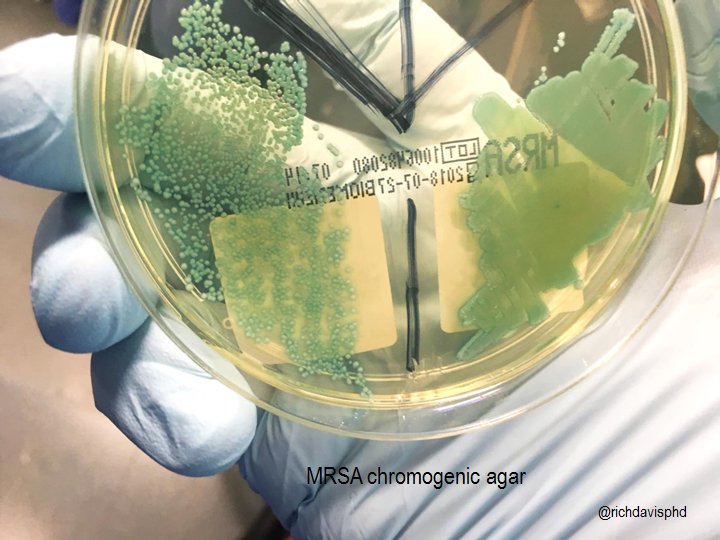
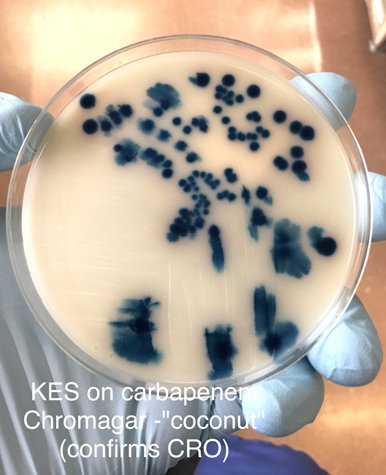
Good for screening test. An agar is impregnated with the antibiotic (e.g. oxacillin - MRSA, ertapenem – Carbapenem Resistance). If an organism grows, it’s likely a resistant isolate.
Instead of being in a disk, the antibiotic is in a strip with high- to low-concentration of the antibiotic. The growth of the bacteria next to the strip can tell you the inhibitory concentration.

A modification of the disk method.
Step 1: incubate a disk w/ a bacteria for 2-4hrs.
Step 2: see if that disk can still inhibit a different bacteria (E. coli) on a plate!
see: twitter.com/search?q=cim%2…
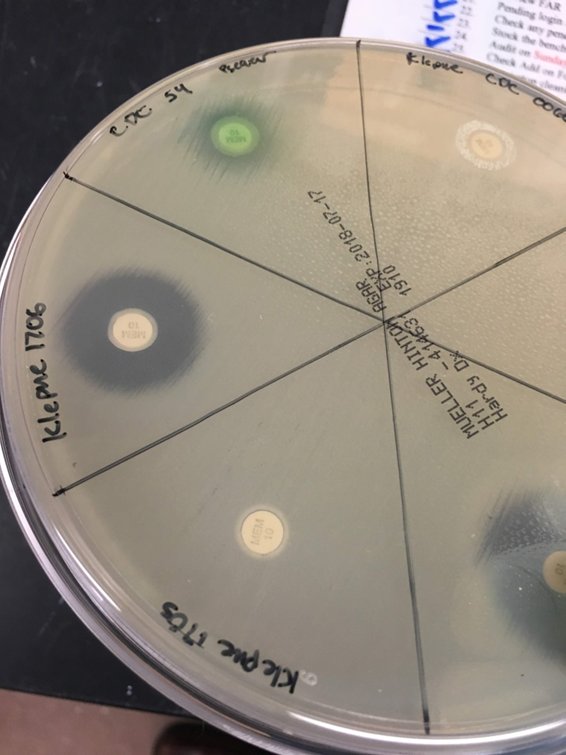
You have a disk (a beta lactam drug alone) and a disk PLUS a beta-lactamASE inhibitor. If inhibitor makes bacteria more susceptible, the beta-lactamase was in the bacteria. see:

But other methods include looking for presence of a protein or gene that PREDICTS resistance.
One benefit: these can be faster! (esp PCR)
Why is MRSA resistant to methycillin/oxacillin? Bc the beta-lactam drug can't bind to the bacteria's mutant penicillin binding protein PBP2a
Lateral flow test: a band forms if PBP2a is present.


This is a HUGE category. MRSA. Carabapenemase. M. tuberculosis resistance genes. There are hundreds of potential resistance mechanism-encoding genes. One benefit: (usually) faster than culture!
One rapid PCR test, Biofire's BCID filmarray, includes some major #AntibioticResistance targets.
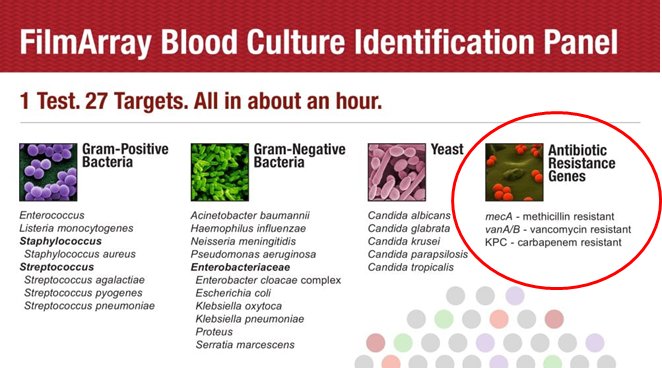

Answers:
1) it can cost a lot!
2) many bacteria have multiple resistance mechanisms
3) in a mixed infection, you can get confounding results!
4) many #AntibioticResistant mechanism genes remain unknown
This was by no means comprehensive. #ASMClinMicro #tweetorial
1) @IDstewardship's website training.idstewardship.com
2) The Sanford Abx guide: sanfordguide.com/products/print…
3) Your nearest ID pharmacist!
#AntibioticResistance
CLSI @CLSI_LabNews M100 manual (accessible for free) is a wealth of information clsi.org/standards/prod…
If there are other antibiotic resources PLEASE comment and I'll add them to this thread!
#ASMClinMicro
That'll do it for me! Thanks for following along!
#tweetorial #ASMClinMicro